
Enhanced expression and purification of nucleotide-specific ribonucleases MC1 and Cusativin - ScienceDirect

Purification of hetero-oligomeric protein variants using a modified tandem affinity purification approach - ScienceDirect

Protein and antibody purification followed by immunoprecipitation of MYB and GATA zinc finger-type maize proteins with magnetic beads - ScienceDirect
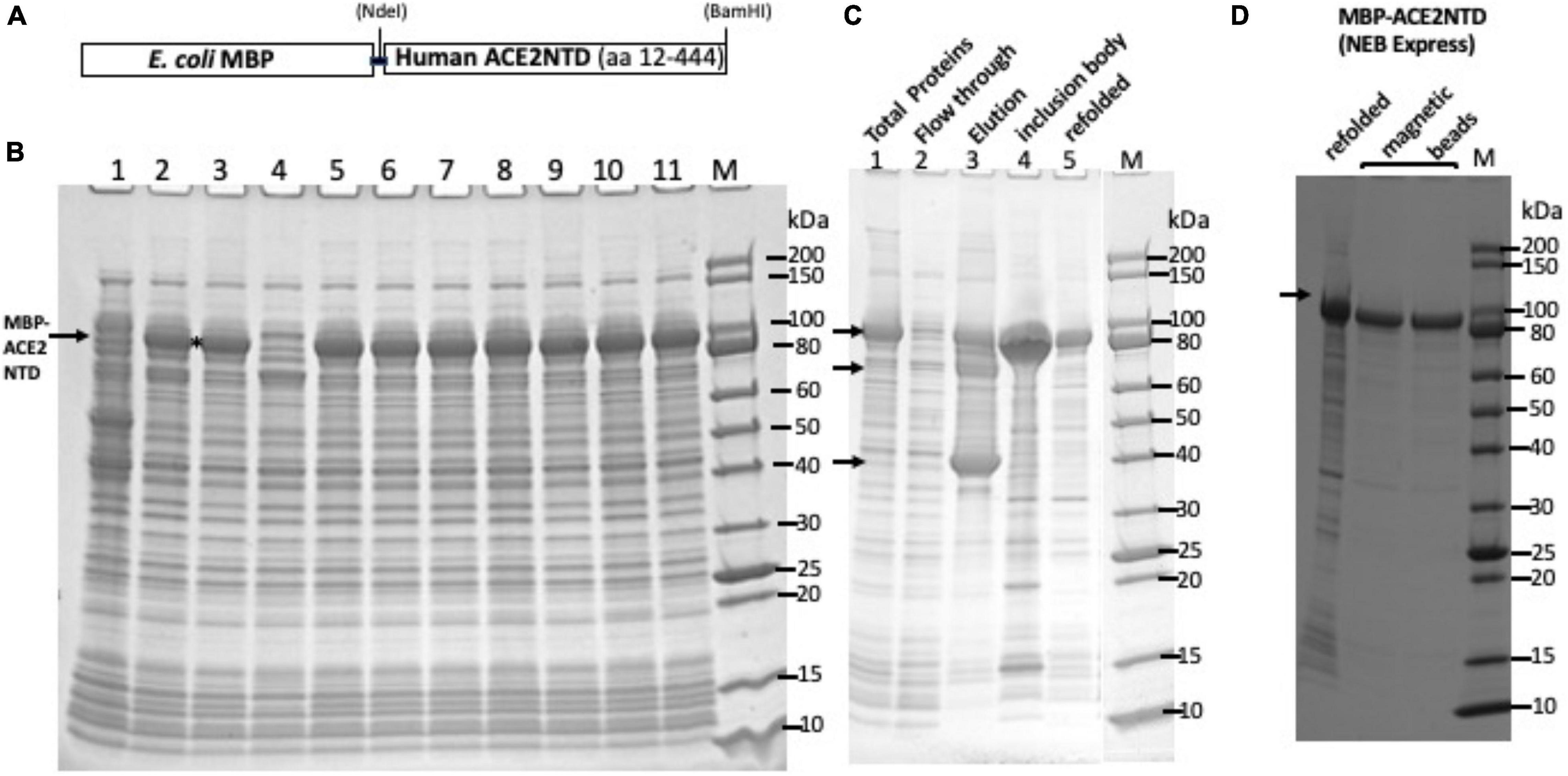
Frontiers | Expression of Human ACE2 N-terminal Domain, Part of the Receptor for SARS-CoV-2, in Fusion With Maltose-Binding Protein, E. coli Ribonuclease I and Human RNase A
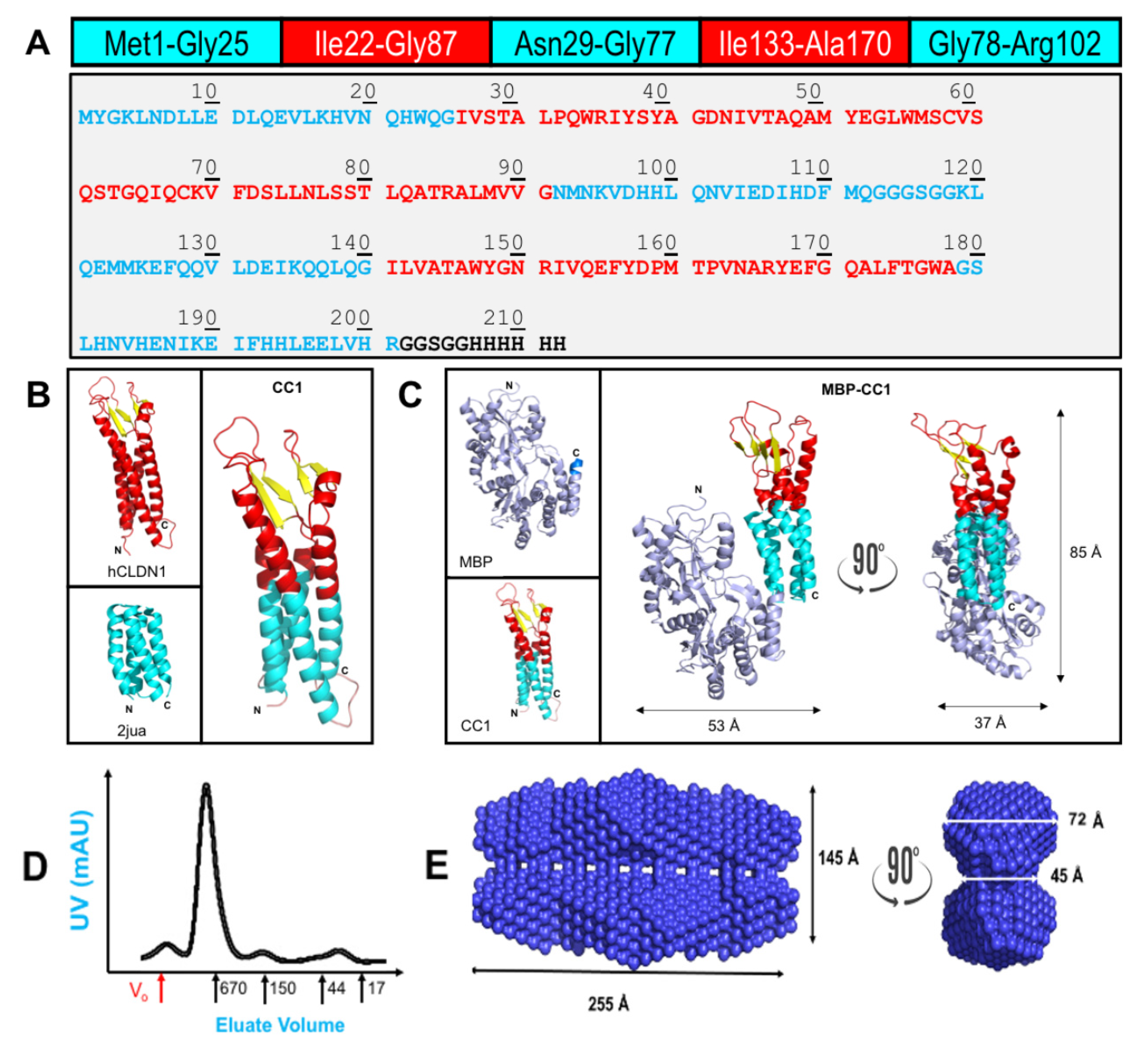
IJMS | Free Full-Text | Chimeric Claudins: A New Tool to Study Tight Junction Structure and Function
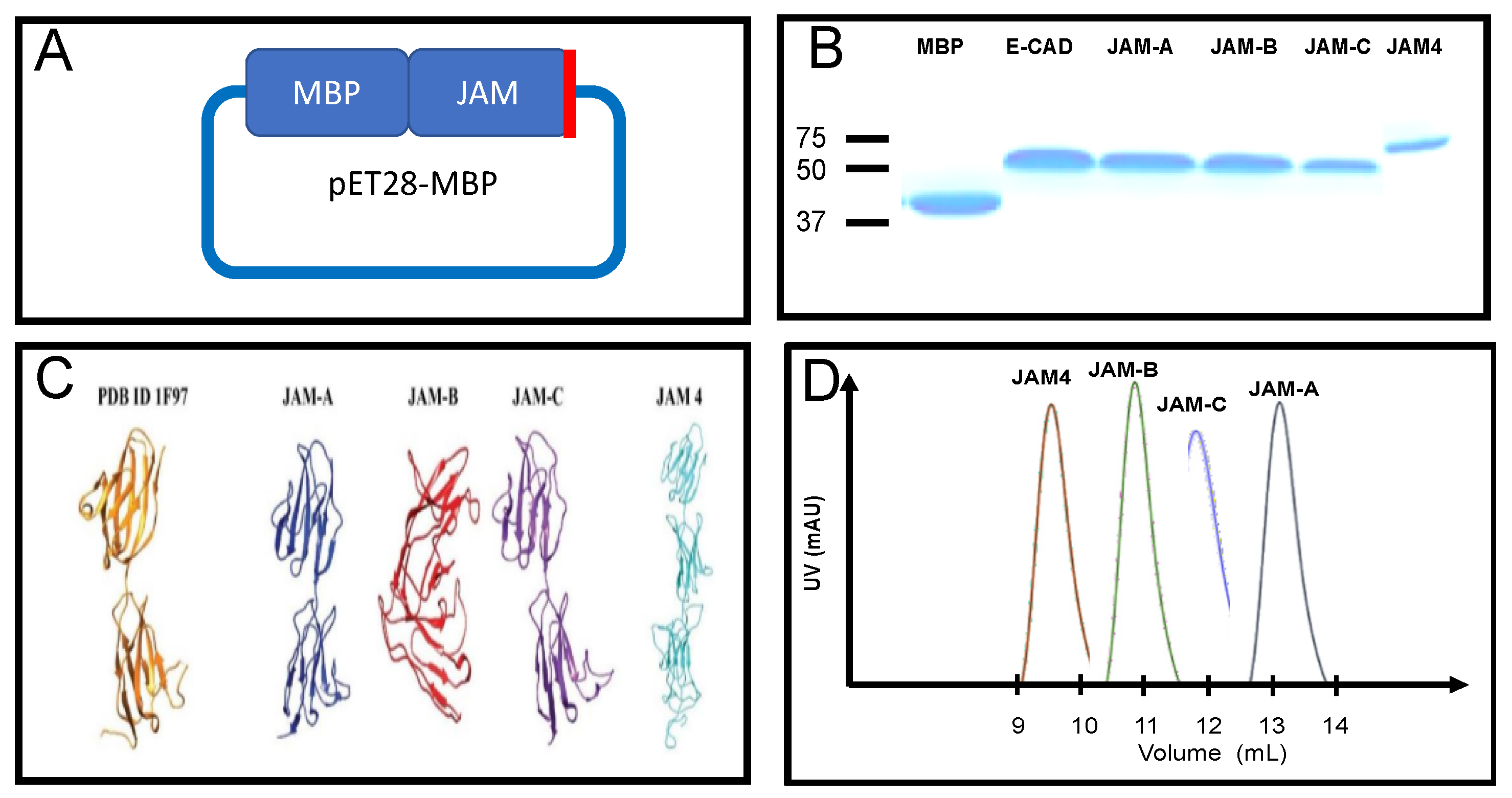
IJMS | Free Full-Text | Molecular Characterization of the Extracellular Domain of Human Junctional Adhesion Proteins

Conserved Characteristics of NMPylation Activities of Alpha- and Betacoronavirus NiRAN Domains | Journal of Virology

Mapping of the regions implicated in nuclear localization of multi‐functional DNA repair endonuclease XPF‐ERCC1 - Akahori - 2022 - Genes to Cells - Wiley Online Library

Expression of human ACE2 N-terminal domain, part of the receptor for SARS-CoV-2, in fusion with maltose binding protein, E. coli ribonuclease I and human RNase A | bioRxiv

Expression of human ACE2 N-terminal domain, part of the receptor for SARS-CoV-2, in fusion with maltose binding protein, E. coli ribonuclease I and human RNase A | bioRxiv
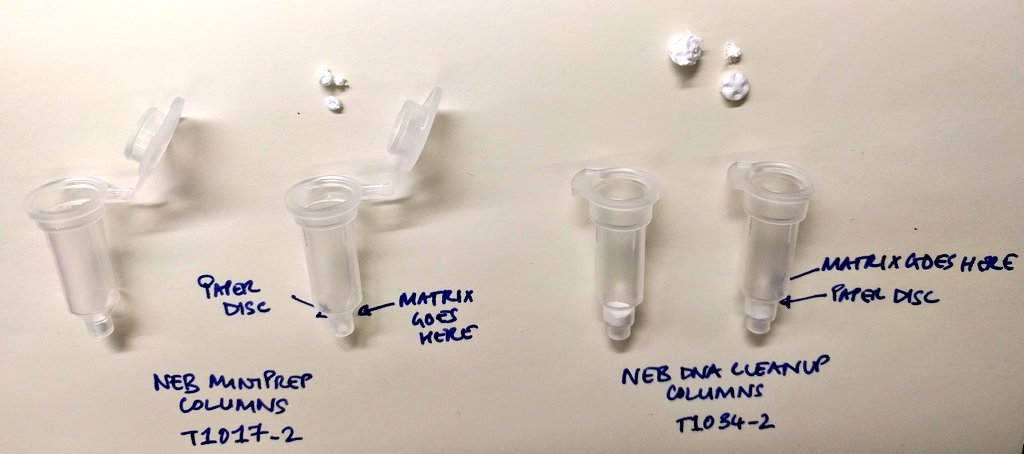


![NEB] NEBExpress® T4 Lysozyme | 코람바이오텍(주)/코람바이오젠(주) NEB] NEBExpress® T4 Lysozyme | 코람바이오텍(주)/코람바이오젠(주)](https://www.korambiotech.com/wp-content/uploads/2023/06/nebexpress-figure1-min.png)
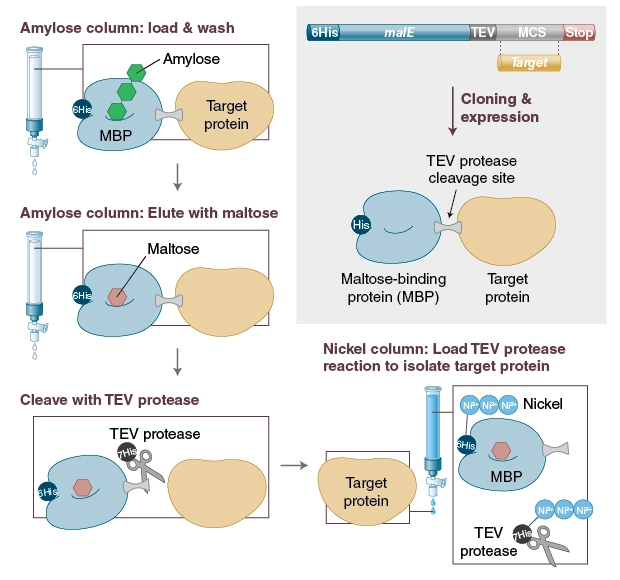





![NEB] NEBExpress® T4 Lysozyme | 코람바이오텍(주)/코람바이오젠(주) NEB] NEBExpress® T4 Lysozyme | 코람바이오텍(주)/코람바이오젠(주)](https://www.korambiotech.com/wp-content/uploads/2023/06/nebexpress-figure2-min.png)

